Statistics
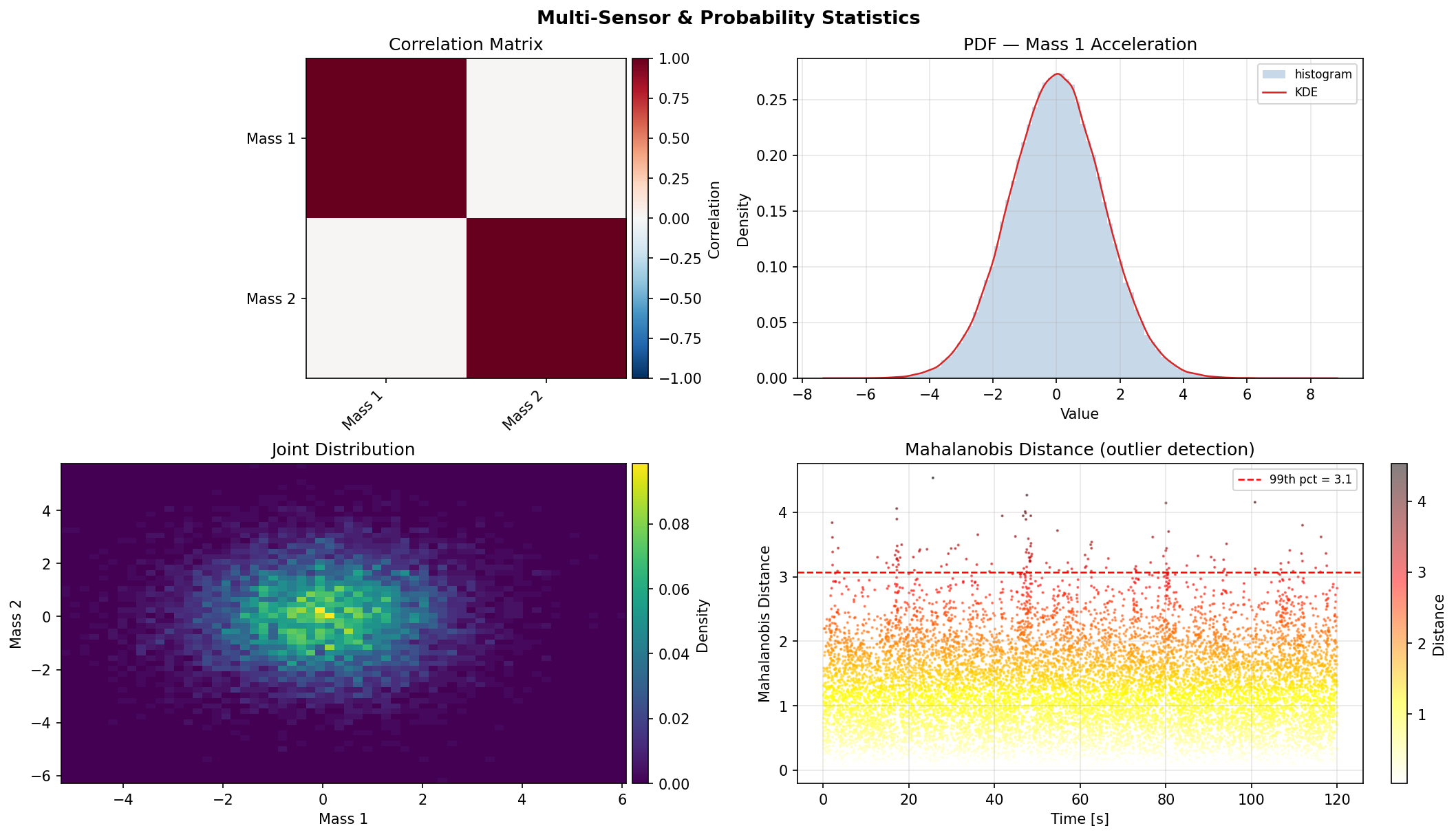
Probability density estimation and joint statistics for signal characterisation: KDE, histograms, joint distributions, covariance, and Mahalanobis distance.
dspkit.statistics.pdf_estimate(x, n_points=256, bandwidth=None)
Kernel density estimate (KDE) of a signal's probability density function.
Uses a Gaussian kernel with automatic or user-specified bandwidth.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
x
|
(array_like, shape(N))
|
Signal samples. |
required |
n_points
|
int
|
Number of evaluation points (default 256). |
256
|
bandwidth
|
float or None
|
KDE bandwidth (standard deviation of the Gaussian kernel).
If |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
xi |
(ndarray, shape(n_points))
|
Evaluation points (range of |
density |
(ndarray, shape(n_points))
|
Estimated PDF values. |
Source code in dspkit/statistics.py
dspkit.statistics.histogram(x, bins=50, density=True)
Normalised histogram (empirical PDF approximation).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
x
|
(array_like, shape(N))
|
Signal samples. |
required |
bins
|
int or array_like
|
Number of bins or bin edges. |
50
|
density
|
bool
|
If |
True
|
Returns:
| Name | Type | Description |
|---|---|---|
bin_centres |
ndarray
|
Centre of each bin. |
counts |
ndarray
|
Histogram values (probability density if |
Source code in dspkit/statistics.py
dspkit.statistics.joint_histogram(x, y, bins=50, density=True)
2D histogram (empirical joint PDF) of two signals.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
x
|
(array_like, shape(N))
|
Signal samples (must have equal length). |
required |
y
|
(array_like, shape(N))
|
Signal samples (must have equal length). |
required |
bins
|
int or (int, int)
|
Number of bins in each dimension. |
50
|
density
|
bool
|
If |
True
|
Returns:
| Name | Type | Description |
|---|---|---|
x_centres |
(ndarray, shape(nx))
|
Bin centres along x. |
y_centres |
(ndarray, shape(ny))
|
Bin centres along y. |
H |
(ndarray, shape(nx, ny))
|
Joint histogram values. |
Source code in dspkit/statistics.py
dspkit.statistics.covariance_matrix(data, bias=False)
Covariance matrix for multi-channel data.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
data
|
(array_like, shape(n_channels, N))
|
Each row is a time series from one sensor. |
required |
bias
|
bool
|
If |
False
|
Returns:
| Name | Type | Description |
|---|---|---|
C |
(ndarray, shape(n_channels, n_channels))
|
Covariance matrix. |
Source code in dspkit/statistics.py
dspkit.statistics.mahalanobis(data, reference=None)
Mahalanobis distance of each time sample from the distribution centre.
Useful for multivariate outlier detection in multi-channel SHM data.
D_M(x) = sqrt( (x - μ)^T · Σ^{-1} · (x - μ) )
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
data
|
(array_like, shape(n_channels, N))
|
Multi-channel time series. Each column is one observation. |
required |
reference
|
(array_like, shape(n_channels, N_ref) or None)
|
Reference data to compute the mean and covariance from.
If |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
distances |
(ndarray, shape(N))
|
Mahalanobis distance of each time sample. |